УДК 636.2:636.082.13:575.174
DIFFERENTIATION OF CATTLE BREEDS WITH THE USE OF MULTILOCUS INTERMICROSATELLITE ANALYSIS (ISSR-PCR)
Yu.A. Stolpovskii1, M. Akhani Azari1, 2, A.N. Evsyukov1, N.V. Kol1, M.N. Ruzina1, L.N. Voronkova1, G.E. Sulimova1
The authors discussed the prospects in application of transgenic technologies for commodity production of fish in commercial aquarium, for the use of fish as bioindikator organism with clear determinate pathological symptoms (for instance, for ecological monitoring, valuation of risk, influence of acute exposure of high doses and chronic effect of small doses of chemical compounds), and also the creation of DNA-vaccines of new generation, raising the safety of fish in aquaculture. The authors marked the necessity of fundamental genetic, biochemical, physiological investigations both mechanisms and consequence of expression of incorporated gene construction, as far as on genetically modified (GM) fish such works are carried out in extremely limited volume. The problem of support of biosafety of GM fish species for natural cenosis was considered. The special attention the author devoted to the application of GM in artificial fish food, that result in the involvement of GM organisms (GMO) in food chain, and to the problems of food security in connection with globalization and monopolization of GMO market.
Keywords: inter simple sequence repeats (ISSR), DNA-markers, breed of cattle, differentiation, phylogenesis.
In the XX century, domestication of cattle (including zebu) was believed to occur in three major centers - India (today’s territory of India and Pakistan), Mediterranean (Mediterranean Sea coast) and South-West Asia (Asia Minor, the Caucasus, Iran) (1). There were no doubts about the common ancestor of modern cattle – aurochs, but no single opinion about place and time of its domestication. Modern methods of DNA analysis helped to clarify this issue. Two centers of domestication were found and two wild ancestors of modern cattle ranked as two different species (2). The first one is located in the Near East between the Black Sea, the Caspian Sea, the Mediterranean Sea and the Persian Gulf; Bos taurus, the ancestor of European cattle was domesticated here. The humped zebu cattle B. indicus originates from the second center (now is Pakistan). The analysis of the D-loop nucleotide sequences of mitochondrial DNA revealed the divergence of wild cattle ancestors dated by 200-1000 thousand years ago, i.e. long before domestication of cattle that occurred 8-10 thousand years ago (2).
Global spreading of domestic animals was greatly connected with human migrations from East to West. Thus, Asians moved to Europe (4-5 thousand years BC) brought there domesticated cattle. Studying the genomic polymorphism of domesticated species using DNA markers can give the arguments to refute or confirm different hypotheses proposed on the basis of archaeological, paleontological, historical and other sciences.
Unfortunately, there are no data about any significant investigations on this issue performed in Eastern Europe and Russia yet. In this regard, the study of gene pools based on ISSR-fingerprinting (ISSR - multilocus estimate of polymorphism of inter simple sequence repeats) is certainly of interest aimed at determination of breeds’ phylogeny and clarifying the origin of modern cattle populations, primarily in the Russian Federation.
The purpose of this research was a comparative study of genetic structure in populations of cattle breeds of European, Asian and African origin, using ISSR analysis.
Technique. The object of study were 761 animals of 19 cattle breeds and 1 selection type (the Caucasian type of Brown Swiss breed from the Republic of Dagestan) represented by two populations - APC "Druzhba» (n = 48) and SUE "Dylymskoe» (n = 56). Among the six studied native breeds, there were two ones created in the former Soviet Union - Bestuzhev (n = 54) and Kostroma (n = 60), and breeds of more ancient origin – Red Estonian (n = 50), Kalmyk (n = 29), Yaroslavl (n = 62), Yakut (n = 30), and Gray Ukrainian steppe cattle (n = 45). The studied foreign breeds were African N'dama (n = 30), New Zealand Friesian (n = 29), Holstein-Friesian (n = 25), Norwegian Red (n = 30), French Norman (n = 23) and Montbeliyarde (n = 29), Indian (B. indicus) Haryana (n = 9), Tharparkar (n = 9) and Sahiwal (n = 7), whose DNA samples were kindly provided by Dr. K. Meade and Dr. D. MacHugh from the UCD College of Agriculture (Dublin, Ireland). DNA samples of German Black-and-White breed (n = 42) were obtained from Dr V. Reiharat and Dr. E. Gernand from the Scientific Research Center for Animal Husbandry of the German Academy Agricultural Sciences (Dummerstorf, Germany). Blood samples of Mongolian Hogorogo (n = 47) and South Gobi cattle (n = 47) were collected during a joint Russian-Mongolian scientific expedition.
For ISSR-PCR, there were used the primers (AG)9C and (GA)9C (3). PCR amplification was performed under the following regime: an initial denaturation - 2 min at 94-95 °C, denaturation – 30 s at 94 °C, annealing – 30 s at 55 °C, synthesis - 2 minutes at 72 ° C, the final synthesis - 7-10 minutes at 72 °C; the number of amplification cycles - 35-37 (the technique was described in detail earlier) (4). The PCR products were fractionated in 2% agarose gel at 120 V for 100-110 min. The molecular weight marker was GeneRulerTM 100 bp DNA Ladder Plus (“MBI Fermentas”, Latvia). After electrophoresis, the gels were stained with ethidium bromide and photographed under shortwave UV light.
The data were statistically processed in Microsoft Office standard programs, multidimensional scaling and principal component analysis – in Statistica 8.0.
Results. A comparative study of cattle breeds of European, Asian and African origin has allowed more fully clarify their phylogenetic relationships. In total, there were revealed 66 PCR products for two types of ISSR-markers, 64 of which were polymorphic. There were synthesized 37 amplification products with (AG)9C primer (including 34 polymorphic) and 29 - with (GA)9C primer (all polymorphic); the length of fragments varied from 2500 to 160 bp. Frequencies of occurrence of the obtained fragments for AG- and GA-ISSR marker systems in 19 breeds and one selection type of cattle are shown in the table.
As a rule, breeds differed from each other by the spectrum of ISSR-fragments and their frequency (Table), as well as by luminescence intensity of particular regions of the fragments on electrophoregram, which apparently reflects different representation of the amplified DNA fragments in cattle genomes.
Thus, for the primer (AG)9C, the largest fragment weighting 2500-2300 bp (A1) was found only in five populations - Mongolian cattle from South Gobi (0,256), Norwegian Red (0,225), Tharparkar (0,184), Haryana (0,118) and Yakut (0,051) breeds. Some of the ISSR-fragments were present in particular breeds with a reliably different frequency (P <0,05). For example, a fragment of 1550-1500 bp (A6) was detected in all breeds (frequency - from 0,408 to 1) except Gray Ukrainian and Kalmyk. In most European breeds, the frequency of the fragment A20 670-640 bp length significantly differed from breeds of Asian origin, particularly from Zebu-like cattle from India (B. indicus). A similar distribution of the fragments was observed using the primer (GA)9C (Table). The obtained results suggest that identified similarities and differences in distribution of ISSR-fragments allow to differentiate the spectra of PCR products and particular cattle breeds as well.
It’s notable that spectra of amplicons obtained with (GA)9C primer were more conservative. In this case, there were found breed-specific fragments and more precise separation of cattle populations by breeding and geographic criteria. For example, the fragment A20 was present only in Red Estonian breed, while A24 – only in Tharparkar. In other words, the primer (GA)9C allows to reveal the polymorphism reflecting interbreed and interpopulation separation/affinity, while the primer (AG)9C helps to perform a more detailed clustering and analysis, that is, to investigate their interpopulation polymorphism. This fact was supported by the established diversity indices: the proportion of interpopulation (interbreed) diversity in total for all studied populations (GST) for the primer (GA)9C amounted to 69,9%, for (AG)9C – 46,2%, inbreeding diversity (HS) equaled 0,0343 and 0,0916, respectively, total diversity (HT) - 0,1139 and 0,1704.
Multidimensional scaling of data and principal component analysis revealed detailed genetic relationships between breeds and made possible the graphical visualization of spatial distribution of the analyzed cattle groups (Fig. 1, 2). It should be emphasized that interpretation of the presented data on intra- and interbreed diversity can alter depending on number of studied breeds.
The history of selection, classification and breeding areas are known for most of the studied breeds. Thus, the breed Haryana, Tharparkar and Sahiwal, as well as N'dama belong to the Afro-Asian group descended from B. indicus; most of other breeds belong to the species B. taurus, while some researchers consider Mongolian, Kalmyk and Yakut breeds as the separate subspecies of B. taurus turano-mongolicus (5, 6). In the analyzed populations, there were the breeds classified by FAO as transboundary international breeds (bred in many countries, such as Holstein-Friesian, Black-and-White), transboundary regional (bred in several countries of one region – eg. N'dama, Red Estonian) and local (most of the studied breeds - Kalmyk, Yaroslavl, Kostroma, etc.).
The frequency of DNA fragments obtained by PCR-amplification with primers (AG)9C (AG-ISSR markers) and (GA)9C (GA-ISSR markers) in populations of native and foreign cattle breed |
|||||||||||||||||||||||
Fragment |
Length, bp |
Bestuzhev |
Brown Swiss |
Brown Swiss |
Holstein-Friesian |
Kalmyk (n = 29) |
Kostroma (n = 60) |
Red Estonian |
South Gobi cattle |
Montbeliyarde (n = 29) |
N’dama (n = 30) |
Norwegian Red |
Norman (n = 23) |
Sahiwal (n = 7) |
Gray Ukrainian |
Tharparkar (n = 9) |
Friesian (n = 29) |
Haryana (n= 9) |
Hogorogo (n = 47) |
German Black-and-White (n = 42) |
Yakut (n =30) |
Yaroslavl (n = 62) |
Total (N = 761) |
A G - I S S R markers |
|||||||||||||||||||||||
А1 |
2500-2300 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,256 |
0,000 |
0,000 |
0,225 |
0,000 |
0,000 |
0,000 |
0,184 |
0,000 |
0,118 |
0,000 |
0,000 |
0,051 |
0,000 |
0,030 |
А2 |
2100-2000 |
0,000 |
0,000 |
1 |
0,368 |
0,000 |
0,000 |
0,000 |
0,044 |
0,475 |
0,247 |
0,000 |
0,489 |
0,155 |
0,000 |
0,000 |
0,090 |
0,057 |
0,242 |
0,074 |
0,087 |
0,418 |
0,193 |
А3 |
1900-1800 |
0,529 |
0,011 |
1 |
0,000 |
0,000 |
0,000 |
0,000 |
0,137 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,184 |
0,000 |
0,012 |
0,144 |
0,120 |
0,139 |
А4 |
1750-1700 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
1 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,008 |
0,066 |
А5 |
1650-1600 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,022 |
0,000 |
0,000 |
0,000 |
0,001 |
А6 |
1550-1500 |
0,764 |
0,567 |
1 |
0,800 |
0,000 |
0,408 |
1 |
1 |
0,737 |
0,817 |
0,684 |
0,792 |
0,622 |
0,000 |
1 |
0,629 |
1 |
0,794 |
0,537 |
1 |
0,820 |
0,699 |
А7 |
1450-1400 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,078 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,011 |
0,000 |
0,000 |
0,000 |
0,005 |
А8 |
1350-1300 |
0,000 |
0,000 |
0,000 |
0,800 |
0,000 |
0,000 |
0,000 |
0,000 |
0,737 |
0,742 |
0,395 |
0,792 |
0,622 |
0,000 |
1 |
0,545 |
1 |
0,000 |
0,000 |
1 |
0,780 |
0,276 |
А9 |
1290-1240 |
0,491 |
0,796 |
0,599 |
0,800 |
0,545 |
0,742 |
1 |
1 |
0,737 |
0,000 |
0,635 |
0,705 |
0,622 |
0,000 |
1 |
0,629 |
1 |
0,516 |
0,345 |
1 |
0,873 |
0,651 |
А10 |
1230-1180 |
0,864 |
0,011 |
0,084 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,017 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,229 |
0,000 |
0,000 |
0,081 |
А11 |
1170-1120 |
0,000 |
0,021 |
0,599 |
0,000 |
0,000 |
0,000 |
0,000 |
0,089 |
0,035 |
0,684 |
0,000 |
0,000 |
0,000 |
0,057 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,083 |
А12 |
1110-1060 |
0,000 |
0,032 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,022 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,087 |
0,000 |
0,006 |
А13 |
1050-1000 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
0,792 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
0,994 |
А14 |
990-940 |
0,000 |
0,011 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,017 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,001 |
А15 |
930-880 |
0,667 |
0,000 |
0,000 |
0,800 |
0,000 |
0,000 |
0,000 |
0,000 |
0,629 |
0,635 |
0,452 |
0,639 |
0,622 |
0,000 |
1 |
0,585 |
1 |
0,000 |
0,488 |
1 |
1 |
0,359 |
А16 |
870-820 |
0,808 |
1 |
1 |
0,800 |
1 |
1 |
1 |
1 |
0,585 |
0,635 |
0,423 |
0,583 |
0,622 |
1 |
1 |
0,629 |
1 |
0,794 |
0,488 |
1 |
1 |
0,856 |
А17 |
810-760 |
0,808 |
0,796 |
1 |
0,800 |
0,281 |
0,317 |
1 |
1 |
0,678 |
0,635 |
0,635 |
0,583 |
0,622 |
0,000 |
1 |
0,678 |
1 |
0,474 |
0,733 |
1 |
1 |
0,710 |
А18 |
750-720 |
0,000 |
0,043 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,017 |
0,000 |
0,000 |
0,285 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,020 |
А19 |
710-680 |
0,067 |
0,592 |
0,418 |
0,062 |
0,000 |
0,043 |
0,000 |
0,435 |
0,129 |
0,000 |
0,204 |
0,045 |
0,622 |
0,046 |
1 |
0,035 |
0,000 |
0,113 |
0,000 |
0,553 |
0,016 |
0,171 |
А20 |
670-640 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
0,678 |
1 |
0,684 |
0,792 |
0,622 |
1 |
0,529 |
1 |
0,667 |
0,854 |
1 |
1 |
1 |
|
А21 |
630-600 |
0,000 |
0,032 |
0,000 |
0,000 |
0,000 |
0,008 |
0,062 |
0,000 |
0,017 |
0,000 |
0,106 |
0,000 |
0,000 |
0,081 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,016 |
А22 |
590-560 |
0,009 |
0,371 |
0,074 |
0,000 |
0,000 |
0,000 |
0,000 |
0,044 |
0,000 |
0,000 |
0,034 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,529 |
0,000 |
0,000 |
0,000 |
0,000 |
0,040 |
А23 |
550-530 |
0,019 |
0,011 |
0,000 |
0,000 |
0,000 |
0,008 |
0,000 |
0,000 |
0,017 |
0,000 |
0,163 |
0,045 |
0,244 |
0,000 |
0,184 |
0,000 |
0,333 |
0,011 |
0,012 |
0,163 |
0,000 |
0,027 |
А24 |
520-500 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
А25 |
490-470 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
0,792 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
0,994 |
А26 |
460-440 |
0,009 |
0,000 |
1 |
0,000 |
0,000 |
0,000 |
0,010 |
0,000 |
0,000 |
0,000 |
0,017 |
0,000 |
0,000 |
0,000 |
0,057 |
0,000 |
0,000 |
0,022 |
0,000 |
0,000 |
0,000 |
0,078 |
А27 |
430-410 |
0,047 |
1 |
0,866 |
0,062 |
0,000 |
0,817 |
0,000 |
1 |
0,035 |
0,000 |
0,051 |
0,000 |
0,074 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,263 |
А28 |
400-380 |
1 |
0,711 |
0,000 |
1 |
1 |
0,742 |
0,800 |
1 |
0,814 |
1 |
1 |
1 |
0,244 |
1 |
0,333 |
1 |
0,423 |
0,674 |
1 |
1 |
1 |
0,826 |
А29 |
370-360 |
0,391 |
0,856 |
0,176 |
0,717 |
0,212 |
1 |
0,368 |
0,348 |
0,357 |
0,395 |
0,684 |
0,192 |
0,000 |
1 |
0,000 |
0,545 |
0,000 |
0,364 |
0,244 |
0,553 |
0,508 |
0,492 |
А30 |
350-340 |
0,000 |
0,000 |
0,000 |
0,000 |
0,509 |
0,000 |
0,717 |
0,000 |
0,000 |
0,000 |
0,225 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,075 |
А31 |
330-320 |
0,000 |
0,146 |
0,055 |
0,084 |
0,053 |
0,215 |
0,010 |
0,101 |
0,072 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,197 |
0,061 |
А32 |
310-300 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,008 |
0,010 |
0,000 |
0,000 |
0,000 |
0,051 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,003 |
А33 |
290-280 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
А34 |
270-260 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,057 |
0,000 |
0,000 |
0,000 |
0,000 |
0,001 |
А35 |
250-240 |
0,549 |
1 |
1 |
1 |
0,814 |
0,613 |
0,755 |
1 |
0,585 |
1 |
0,817 |
0,341 |
0,622 |
0,529 |
1 |
0,737 |
1 |
0,101 |
0,733 |
0,742 |
0,716 |
0,726 |
А36 |
230-220 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
А37 |
210-200 |
0,009 |
0,223 |
1 |
0,000 |
0,000 |
0,270 |
0,000 |
1 |
0,090 |
0,069 |
0,124 |
0,022 |
0,466 |
0,000 |
0,423 |
0,000 |
0,529 |
0,000 |
0,260 |
0,423 |
0,158 |
0,242 |
А38 |
180-160 |
0,000 |
0,171 |
0,000 |
0,000 |
0,000 |
0,069 |
0,000 |
0,000 |
0,000 |
0,000 |
0,087 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,032 |
0,000 |
0,000 |
0,000 |
0,022 |
G A - I S S R markers |
|||||||||||||||||||||||
А1 |
2500-2300 |
0,070 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
1 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,099 |
0,177 |
0,089 |
А2 |
2100-2000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,031 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,002 |
А3 |
1900-1800 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
А4 |
1750-1700 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
А5 |
1650-1600 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
А6 |
1550-1500 |
0,861 |
1 |
1 |
1 |
0,000 |
1 |
1 |
1 |
1 |
1 |
0,804 |
0,517 |
1 |
1 |
1 |
1 |
1 |
0,856 |
0,586 |
0,694 |
1 |
0,889 |
А7 |
1450-1400 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
А8 |
1350-1300 |
0,000 |
0,192 |
0,131 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,087 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,026 |
А9 |
1290-1240 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
А10 |
1230-1180 |
0,760 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
0,804 |
0,484 |
1 |
1 |
0,368 |
1 |
1 |
1 |
1 |
1 |
1 |
0,949 |
А11 |
1170-1120 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
А12 |
1110-1060 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,032 |
0,000 |
0,000 |
0,000 |
0,002 |
А13 |
1050-1000 |
0,804 |
0,857 |
1 |
1 |
0,570 |
1 |
1 |
1 |
1 |
1 |
0,804 |
0,517 |
1 |
1 |
1 |
1 |
1 |
0,000 |
0,586 |
1 |
1 |
0,855 |
А14 |
990-940 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
А15 |
930-880 |
0,000 |
1 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,008 |
0,065 |
А16 |
870-820 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,009 |
0,000 |
0,000 |
0,000 |
0,017 |
0,145 |
0,000 |
0,000 |
0,000 |
0,051 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,016 |
0,008 |
А17 |
810-760 |
0,000 |
1 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
1 |
0,234 |
0,000 |
0,266 |
0,069 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,172 |
0,171 |
0,023 |
0,163 |
А18 |
750-720 |
0,000 |
0,000 |
0,000 |
0,684 |
0,000 |
0,009 |
0,041 |
0,000 |
0,281 |
0,184 |
0,000 |
0,368 |
1 |
1 |
1 |
1 |
0,452 |
0,011 |
0,303 |
0,414 |
0,570 |
0,267 |
А19 |
710-680 |
0,000 |
0,000 |
0,009 |
0,000 |
0,000 |
0,000 |
0,000 |
0,399 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,025 |
Fragment |
Length, bp |
Bestuzhev |
Brown Swiss |
Brown Swiss |
Holstein-Friesian |
Kalmyk (n = 29) |
Kostroma (n = 60) |
Red Estonian |
South Gobi cattle |
Montbeliyarde (n = 29) |
N’dama (n = 30) |
Norwegian Red |
Norman (n = 23) |
Sahiwal (n = 7) |
Gray Ukrainian |
Tharparkar (n = 9) |
Friesian (n = 29) |
Haryana (n= 9) |
Hogorogo (n = 47) |
German Black-and-White (n = 42) |
Yakut (n =30) |
Yaroslavl (n = 62) |
Total (N = 761) |
A G - I S S R markers |
|||||||||||||||||||||||
А20 |
670-640 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
1 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,065 |
А21 |
630-600 |
0,000 |
1 |
1 |
0,106 |
1 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,034 |
0,051 |
1 |
0,172 |
0,016 |
0,000 |
0,251 |
А22 |
590-560 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
А23 |
550-530 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
0,684 |
1 |
1 |
0,000 |
1 |
1 |
1 |
1 |
1 |
1 |
0,975 |
А24 |
520-500 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
1 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,013 |
А25 |
490-470 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
0,817 |
1 |
1 |
0,000 |
1 |
1 |
1 |
1 |
1 |
1 |
0,980 |
А26 |
460-440 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,019 |
0,000 |
0,000 |
0,000 |
1 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,014 |
А27 |
430-410 |
0,000 |
0,000 |
0,000 |
0,017 |
0,000 |
0,000 |
0,225 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,065 |
0,000 |
0,000 |
0,000 |
0,019 |
А28 |
400-380 |
0,000 |
1 |
1 |
0,742 |
0,000 |
0,869 |
0,000 |
0,516 |
0,443 |
0,000 |
0,380 |
0,452 |
1 |
0,000 |
0,293 |
1 |
0,000 |
0,000 |
0,138 |
0,000 |
0,089 |
0,378 |
А29 |
370-360 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,017 |
0,000 |
0,000 |
0,034 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,002 |
А30 |
350-340 |
0,000 |
1 |
0,813 |
1 |
1 |
1 |
0,000 |
1 |
0,585 |
1 |
0,380 |
1 |
1 |
1 |
0,368 |
1 |
0,051 |
0,856 |
0,831 |
1 |
1 |
0,779 |
А31 |
330-320 |
0,010 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,001 |
А32 |
310-300 |
0,000 |
0,000 |
0,000 |
0,087 |
0,000 |
0,000 |
0,000 |
0,000 |
0,331 |
0,000 |
0,123 |
0,106 |
0,074 |
1 |
0,051 |
0,069 |
0,000 |
0,000 |
0,324 |
0,293 |
0,216 |
0,131 |
А33 |
290-280 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,435 |
0,000 |
0,000 |
0,000 |
0,184 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,034 |
А34 |
270-260 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,017 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,001 |
А35 |
250-240 |
0,520 |
1 |
1 |
1 |
1 |
1 |
0,000 |
0,854 |
1 |
0,742 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
0,883 |
А36 |
230-220 |
0,412 |
1 |
1 |
1 |
0,000 |
1 |
0,000 |
1 |
1 |
1 |
0,660 |
1 |
1 |
0,000 |
1 |
1 |
1 |
0,567 |
1 |
1 |
1 |
0,763 |
А37 |
210-200 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
А38 |
180-160 |
0,000 |
1 |
1 |
0,000 |
0,000 |
0,000 |
0,000 |
1 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,000 |
0,200 |
Note. Description of studied cattle breeds – see Technique. |
|||||||||||||||||||||||
|
|
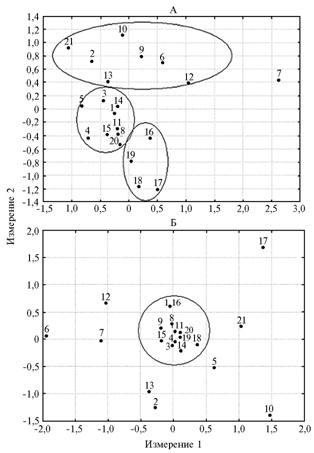
Fig.1. Results of multidimentional scaling on plane performed upon the data obtained for the primers (AG)9C (AG-ISSR markers, A) and (GA)9C (GA-ISSR markers, B): 1-21 — respectively, cattle breeds (populations) Friesian, Kalmyk, Black-and-White, N’dama, Bestuzhev, Brown Swiss (APC “Druzhba”, the Republic of Dagestan), Brown Swiss (SUE “Dylymskoe”, the Republic of Dagestan), Holstein-Friesian, Kostroma, Red Estonian, Montbeliarde, Mongolian South Gobi cattle, Mongolian Hogorogo, Norwegian Red, Norman, Sahival, Tharparkar, Haryana, Yakut, Yaroslavl, Gray Ukrainian. Note: |
|
|
|
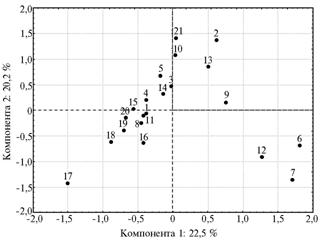
Fig. 2. Spatial distribution of studied populations – native and foreign cattle breeds - relative the principal components for the general set of markers obtained using the primers (AG)9C and (GA)9C: 1-21 — respectively, cattle breeds (populations) Friesian, Kalmyk, Black-and-White, N’dama, Bestuzhev, Brown Swiss (APC “Druzhba”, the Republic of Dagestan), Brown Swiss (SUE “Dylymskoe”, the Republic of Dagestan), Holstein-Friesian, Kostroma, Red Estonian, Montbeliarde, Mongolian South Gobi cattle, Mongolian Hogorogo, Norwegian Red, Norman, Sahival, Tharparkar, Haryana, Yakut, Yaroslavl, Gray Ukrainian. Note: |
|
Local breeds are of great interest in terms of studying the evolution of genus Bos, since many of them carry ancestral form of gene complexes (associations) preserved in their gene pools.
Using the multidimensional scaling method has revealed a different distribution of breeds for AG-ISSR and GA-ISSR markers (Fig. 1). The variability of AG-ISSR markers differentiated cattle population into three groups: major (includes most of the breeds - “a central cloud" located in the central part of the picture), second ("upper cloud") including populations of two Mongolian breeds, Cray Ukrainian, Kalmyk, Estonian Red, Kostroma and the Caucasian type of Brown Swiss breed (SPK "Druzhba") and the third group (the "lower cloud") formed by Sahiwal, Tharparkar, Haryana and Yakut cattle. Such distribution probably reflects a certain phylogenetic relationship or, conversely, the absence of consanguinity between these breeds. The main core was formed by breeds of European origin mainly descent from the Black-and-White root; the separate group of breeds originated from India, Mongolia, breeds of Brown and Red root, Yakut cattle and steppe breeds (Kalmyk and Gray Ukrainian). An isolated location of the Caucasian type of Brown Swiss breed (SUE "Dylymskoe") (Fig. 1) indicates a significant difference of its gene pool (apparently, it was largely represented by the local gene pool of Dagestan cattle) from all other breeds studied in this work.
Interestingly that Yakut cattle representing the most northern part of natural habitat of B. taurus was found to be close to Indian breeds in the scheme. This result was obtained by multidimensional scaling at different conditions (components), and in most cases Haryana, Tharparkar and Yakut cattle were located next to each other. This fact allows two assumptions: first - that Yakut breed many times passed through the so-called "bottleneck" when genetic drift randomly significantly changed the frequency of ISSR-markers, the second - Yakut’s ancestors descended from an ancestor of zebu, and not from Mongolian cattle.
For GA-ISSR markers, the studied breeds showed more uniform distribution reflecting both their kinship and its absence. In general, there was a single pool of breeds and several ones located distant from it – Tharparkar (India), Red Estonian (Russia), Kalmyk (Russia), Grey Ukrainian steppe cattle (Russia), Hogorogo and South Gobi cattle (Mongolia), along with the Caucasian type of Brown Swiss breed. In this study, these gene pools corresponded to edge populations whose breeding history was minimally connected to majority of the studied breeds.
Formation of groups was most adequate to historical events and zootechnical practice, which indicates the prospects for using multi-locus markers in studying phylogeny of cattle. Since the mixed origin of many studied populations, they appeared close to each other at the multidimensional scaling (Fig. 1). Holstein-Friesian and Black-and-White breeds are the connecting links between many breeds, because their gene pools are actively used in interbreed crosses in the territory of former Soviet Union and in the world.
The generalized analysis of interrelations between studied breeds was performed upon the data on both ISSR-markers (AG and GA) resulting in plane projection of principal components (Fig. 2). The observed distribution largely confirmed the results of multidimensional scaling. Most of the breeds were compactly located, eight breeds were located separately from this core and formed three groups: the first - Tharparkar, the second - Mongolian Hogorogo, Gray Ukrainian, Kalmyk, Estonian Red and Kostroma breed, and the third group - two Caucasian populations of Brown Swiss breed and Mongolian cattle from South Gobi. Tharparkar, Gray steppe cattle and Caucasian type of Brown Swiss took the most distant positions, therefore, their gene pools differ from the majority of cattle populations - predominantly European breeds. It can be assumed that these three populations descend not from a single European ancestor, but from two Asian ancestral species or they have polyphyletic origin.
Thus, the authors have established similarities and differences of ISSR-fragments’ frequencies in several cattle breeds of different origin, which allows to differentiate the species Bos taurus and B. indicus. Differentiation and characteristics of genetic relationships between breeds based on ISSR-fingerprinting (including the authors’ findings) can be used in planning selection strategies (eg, interbreeding) and in further studies of phylogeny of B. taurus and B. indicus. Moreover, since the results of ISSR-labeling accurately reflect history of breeds’ selection, there’s an opportunity to use this technique for studying the evolution not only in the genus Bos, but in the species Homo sapiens as well. The data obtained in comparative studies of entire genomes or individual DNA fragments of farm animals can help the reconstruction of human settlement on Earth, the formation of populations from different ethnic groups, as well as the emergence of cultural and technological innovations, such as arise and spread of livestock breeding in the world.
The authors thank Tsendsuren Tsedev (Mongolia) and G.S. Karaev (the Dagestan Union of Livestock Breeding, Russia) for the provided opportunity of obtaining cattle blood samples.
REFERENCES
1. Krasota V.F., Lobanov V.T. and Dzhaparidze T.G., Razvedenie sel’skokhozyaistvennykh zhivotnykh (Breeding Farm Animals), Moscow, 1983.
2. Loftus R.T., MacHugh D.E., Bradley D.G. et al, Evidence for Two Independent Domestications of Cattle, PNAS US, 1994, vol. 91, pp. 2757-2761.
3. Zietkiewicz E., Rafalski A. and Labuda D., Genome Fingerprinting by Sequence Repeat (SSR)-Anchored Polymerase Chain Reaction Amplification, Genomics, 1994, vol. 20, pp. 176-183.
4. Stolpovskii Yu.A. Shmiit L.V., Kol N.V., Evsukov A.N. Ruzina M.N., Churgui-ool O.I. and Sulimova G.E., Analysis of Genetic Variability and Phylogenetic Relations in Populations of Tuvinian Short-Fat-Tailed Seep with ISSR-Markers, S.-kh. biol., 2009, no. 6, pp. 34-43.
5. Kolesnik N.N., Jevolutsiya krupnogo rogatogo skota (Evolution of Cattle), Stalinabad, 1949.
6. Dmitriev N.G., Porody skota po stranam mira. Spravochnaya kniga (Cattle Breeds of the World: Reference Book), Leningrad, 1978.
1N.I. Vavilov Institute of General Genetics, RAAS, Moscow119991, Russia, |
Received February 16, 2010
|